Authors: Karanpartap Singh, Adam Turnbull, Mohammad Abbasi, Kilian Pohl, Feng Vankee Lin, Ehsan Adeli
arXiv Preprint (Submitted to MICCAI 2026).
Abstract: Understanding how large-scale functional brain networks reorganize during cognitive decline remains a central challenge in neuroimaging. While recent self-supervised models have shown promise for learning representations from resting-state fMRI, their internal mechanisms are difficult to interpret, limiting mechanistic insight. We propose BrainInterNet, a network-aware self-supervised framework based on masked reconstruction with cross-attention that explicitly models inter-network dependencies in rs-fMRI. By selectively masking predefined functional networks and reconstructing them from remaining context, our approach enables direct quantification of network predictability and interpretable analysis of cross-network interactions. We train BrainInterNet on multi-cohort fMRI data (from the ABCD, HCP Development, HCP Young Adults, and HCP Aging datasets) and evaluate on the Alzheimer's Disease Neuroimaging Initiative (ADNI) dataset, in total comprising 5,582 recordings. Our method reveals systematic alterations in the brain's network interactions under AD, including in the default mode, limbic, and attention networks. In parallel, the learned representations support accurate Alzheimer's-spectrum classification and yield a compact summary marker that tracks disease severity longitudinally. Together, these results demonstrate that network-guided masked modeling with cross-attention provides an interpretable and effective framework for characterizing functional reorganization in neurodegeneration.
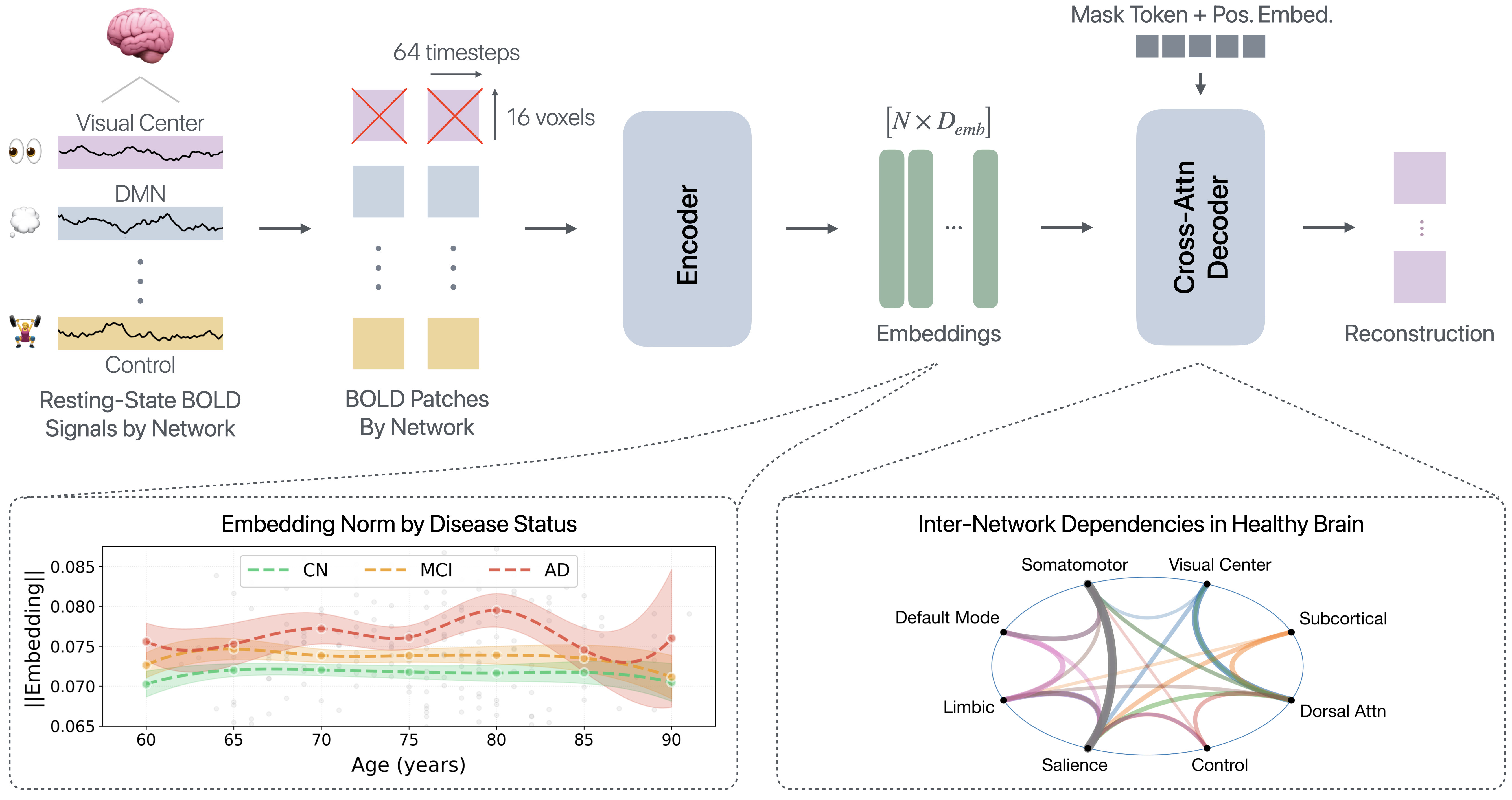
Overview of the proposed framework for modeling inter-network interactions in resting-state fMRI. Resting-state BOLD signals are parcellated using the DiFuMo atlas and divided into non-overlapping temporal-spatial patches. Patches are grouped by Yeo functional networks [Yeo et al., 2011], and those belonging to a selected target network are masked. The encoder processes unmasked tokens to produce contextualized embeddings, while a cross-attention decoder reconstructs the masked network using the context and position embeddings. Learned embeddings reveal separation by disease status, offering a simple metric for mapping neurodegeneration trajectories (bottom left). Meanwhile, the cross-attention decoder provides interpretability into how predictions are made, offering a proxy for inter-network brain dependencies (bottom right).
Authors: Karanpartap Singh, Neil Band, Ehsan Adeli
arXiv Preprint (Submitted to ICML 2026).
Abstract: As the cost of pretraining large language models grows, there is continued interest in strategies to improve learning efficiency during this core training stage. Motivated by cognitive development, where humans gradually build knowledge as their brains mature, we propose Curriculum-Guided Layer Scaling (CGLS), a framework for compute-efficient pretraining that synchronizes increasing data difficulty with model growth through progressive layer stacking (i.e. gradually adding layers during training). At the 100M parameter scale, using a curriculum transitioning from synthetic short stories to general web data, CGLS outperforms baseline methods on the question-answering benchmarks PIQA and ARC. Pretraining at the 1.2B scale, we stratify the DataComp-LM corpus with a DistilBERT-based classifier and progress from general text to highly technical or specialized content. Our results show that progressively increasing model depth alongside sample difficulty leads to better generalization and zero-shot performance on various downstream benchmarks. Altogether, our findings demonstrate that CGLS unlocks the potential of progressive stacking, offering a simple yet effective strategy for improving generalization on knowledge-intensive and reasoning tasks.

Curriculum-Guided Layer Scaling (CGLS) is a new paradigm for compute-efficient language model pretraining that grows data complexity and model depth in tandem. We illustrate CGLS for a Llama-3.2-1B scale model with four training stages. Training begins with an 8-layer model on a data split consisting equally of data from all levels (high-school, undergraduate, and graduate). The learned weights from this stage are transferred to a larger 10-layer model, freezing the pretrained weights and training the new layers on a small, balanced data split for better initialization. The entire model is then unfrozen and pretrained on the more difficult data split. This process is repeated until the target model scale is reached.
Authors: Karanpartap Singh, James Zou
Published @ Transactions on Machine Learning Research (TMLR), 2024, OpenReview.
Abstract: With the increasing use of large-language models (LLMs) like ChatGPT, watermarking has emerged as a promising approach for tracing machine-generated content. However, research on LLM watermarking often relies on simple perplexity or diversity-based measures to assess the quality of watermarked text, which can mask important limitations in watermarking. Here we introduce two new easy-to-use methods for evaluating watermarking algorithms for LLMs: 1) evaluation by LLM-judger with specific guidelines; and 2) binary classification on text embeddings to distinguish between watermarked and unwatermarked text. We apply these methods to characterize the effectiveness of current watermarking techniques. Our experiments, conducted across various datasets, reveal that current watermarking methods are moderately detectable by even simple classifiers, challenging the notion of watermarking subtlety. We also found, through the LLM judger, that watermarking impacts text quality, especially in degrading the coherence and depth of the response. Our findings underscore the trade-off between watermark robustness and text quality and highlight the importance of having more informative metrics to assess watermarking quality.

Can watermarked outputs from large language models be distinguished with a black-box approach? We answer this question through two new methods for evaluating LLM watermarks, showing that independent classifiers and judgers with no prior knowledge of watermarking algorithms prefer or can effectively classify watermarked outputs.
Authors: Kasra Naftchi-Ardebili*, Karanpartap Singh*, Reza Pourabolghasem, Gerald R. Popelka, Kim Butts Pauly (*equal contribution)
Published @ Medical Physics, 2025, Wiley Online Library. Best Talk Award Winner @ ISTU 2023.
Abstract: Transcranial ultrasound stimulation (TUS) has emerged as a promising tool in both clinical and research settings due to its potential to modulate neuronal activity non-invasively. The method delivers focused ultrasound waves to precise regions in the brain, enabling targeted energy deposition. The medical importance of TUS is evidenced by the thirty three ongoing clinical trials, covering conditions such as opioid addiction, Alzheimer's disease, dementia, epilepsy, and glioblastoma. In addition to careful design of ultrasound parameters, treatments with TUS require precise computation of the location and pressure at the focal spot. Heterogeneity of the skull aberrates the incident ultrasound beams, and if uncorrected, poses the risk of off-target sonication or inadequate energy delivery to the neural tissue. For clinical settings, this phase aberration correction must be done within a few seconds. However, physics-informed simulation software suffer from an inherent trade-off between accuracy and efficiency. As such, commercial devices use fast but lower accuracy methods to meet the efficient run times suitable for clinical applications. We present TUSNet, a deep learning approach to address this inherent trade-off between accuracy and efficiency. TUSNet can compute the transcranial ultrasound pressure field within a fraction of a second (1000x faster than k-Wave, a MATLAB-based acoustic simulation package), while achieving over 99% accuracy of the peak pressure at the focal spot with a mean positioning error of 0.1 mm, when compared to a ground truth from k-Wave.

Visual comparison between the TUSNet outputs and the ground truth over two examples. Compared to the ground truth, TUSNet predicts ultrasound pressure fields with nearly identical peak focal pressure, focal spot shape, and reflections inside the skull. Input: Input consists of the transducer elements lined up above the skull, a waveguide, and the target. Background is removed. Ground Truth: k-wave is tasked with simulating the ground truth phase aberration-corrected pressure field using time reversal. TUSNet Output, Absolute Pressure Field: The TUSNet-generated pressure field in Pascals, rather than normalized to some arbitrary maximum value. TUSNet Output, Phase Vector: Phase aberration-corrected pressure field simulated by k-wave based on the TUSNet phase vector rather than time reversal.
Authors: Kasra Naftchi-Ardebili*, Karanpartap Singh*, Reza Pourabolghasem, Pejman Ghanouni, Gerald R. Popelka, Kim Butts Pauly (*equal contribution)
arXiv Preprint (In Review @ Medical Physics)
Abstract: Deep learning offers potential for various healthcare applications involving the human skull but requires extensive datasets of curated medical images. To overcome this challenge, we propose SkullGAN, a generative adversarial network (GAN), to create large datasets of synthetic skull CT slices, reducing reliance on real images and accelerating the integration of machine learning into healthcare. In our method, CT slices of 38 subjects were fed to SkullGAN, a neural network comprising over 200 million parameters. The synthetic skull images generated were evaluated based on three quantitative radiological features: skull density ratio (SDR), mean thickness, and mean intensity. They were further analyzed using t-distributed stochastic neighbor embedding (t-SNE) and by applying the SkullGAN discriminator as a classifier. The results showed that SkullGAN-generated images demonstrated similar key quantitative radiological features to real skulls. Further definitive analysis was undertaken by applying the discriminator of SkullGAN, where the SkullGAN discriminator classified 56.5% of a test set of real skull images and 55.9% of the SkullGAN-generated images as reals (the theoretical optimum being 50%), demonstrating that the SkullGAN-generated skull set is indistinguishable from the real skull set - within the limits of our nonlinear classifier. Therefore, SkullGAN makes it possible to generate large numbers of synthetic skull CT segments, necessary for training neural networks for medical applications involving the human skull. This mitigates challenges associated with preparing large, high-quality training datasets, such as access, capital, time, and the need for domain expertise.
1. A. Stanziola et al., Journal of Computational Physics, vol. 441, p. 110430, 2021.

SkullGAN generator and training pipeline. SkullGAN was first pre-trained on the Celeb-A dataset, and then trained on human skull CTs. In contrast to random initialization of the weights for training on the human skull CTs, pre-training yielded layers with fine-tuned weights for detecting edges and resulted in better quality skull segment images, with finer definition both in contour and interior bone structure.
Authors: Karanpartap Singh, Benjamin G. Hawkins
Abstract: Electrowetting is an electrokinetic effect whereby an applied electric field induces changes in the measured contact angle at a fluid-surface contact line. On hydrophobic, dielectric electrode surfaces, this effect generates droplet motion termed “electrowetting on dielectric” or EWOD. Applications of this phenomenon range from lab-on-a-chip to liquid lenses capable of altering their topology and focus within milliseconds. Electrowetting or EWOD theoretical models quantifying this effect fall into two paradigms: the Young-Lippman and the electromechanical theories. In this work, both paradigms were simulated to predict the velocity of a water droplet moving over an array of electrodes. Results were compared to experimental observations of measured velocities for two dielectric films: ETFE and household cling film. Theoretical model parameters, namely the length scale of the Maxwell force on the droplet, were also determined to align simulation and experiment. The results reveal the trend of droplet velocity in relation to applied voltage, and recapitulate the relationship between the two models.

A) Filmed droplet motion displaying the initial deformation and subsequent motion of the droplet in an open electrowetting configuration. B) Young-Lippman model simulation with only deformation and contact angle change modeled. Colors and color bars represent the velocity fields of the droplets. C) Electromechanical model simulation with only the electromechanical force on the droplet modeled.
Video: some fun footage moving water around purely with electricity / electromagnetic fields, for your viewing pleasure :).
Authors: Karanpartap Singh, Mi Hyun Choi, Gerald Popelka, Kim Butts Pauly
Abstract: Previous studies have demonstrated that transcranial ultrasound stimulation (TUS) leads to varying levels of suppression or excitation of neural activity at the targeted brain region depending on signal parameters – including signal intensity, signal duration, pulse repetition frequency, and pulse duration1-5. However, many studies underreport or inconsistently report these metrics6, preventing the field from converging on a clear relationship between sonication parameters and respective neural effects. To advance TUS as a neuromodulatory tool that can create predictable neural responses, there is a need for systematic testing and reporting of safe and confound-free parameters and brain regions. A safety metric based on FDA guidelines for mechanical and thermal indices7 and an audibility metric for assessing unintended auditory activation confounds in mice were incorporated into a web-based computational tool8, written in Python and hosted through the Flask library. When users either manually input experimental parameters (Figure 1A) or upload hydrophone measurements, the tool provides a standardized report on key TUS parameters (Figure 1B). The report also includes an ideal reconstruction of the inputted signal for confirmation as well as an assessment of the signal’s audibility in mice. This tool is easily accessible to aid in the selection of appropriate, safe, and inaudible signals for neuromodulation and can be used to guide standardized parameter reporting across studies, facilitating reproducibility and inter-study comparisons.
1. S.S. Yoo et al., Neuroimage, vol. 56, p. 1267-1275, 2021. 2. R.L. King et al., Biol., vol. 39, pp. 312-331, 2013. 3. H. Kim et al., Brain Stimul., vol. 7, pp. 748-756, 2014. 4. M. Plaksin et al., Eneuro, vol. 3, 2016. 5. K. Yu et al., IEEE Trans. Biomed., Eng. 63, p. 1787–1794, 2016. 6. C. Pasquinelli et al., Brain stimulation, vol. 12(6), p. 1367–1380, 2019. 7. T. R. Nelson et al., vol. 28(2), p. 139–150, 2009. 8. K. Singh, sr.karanps.com, 2021.
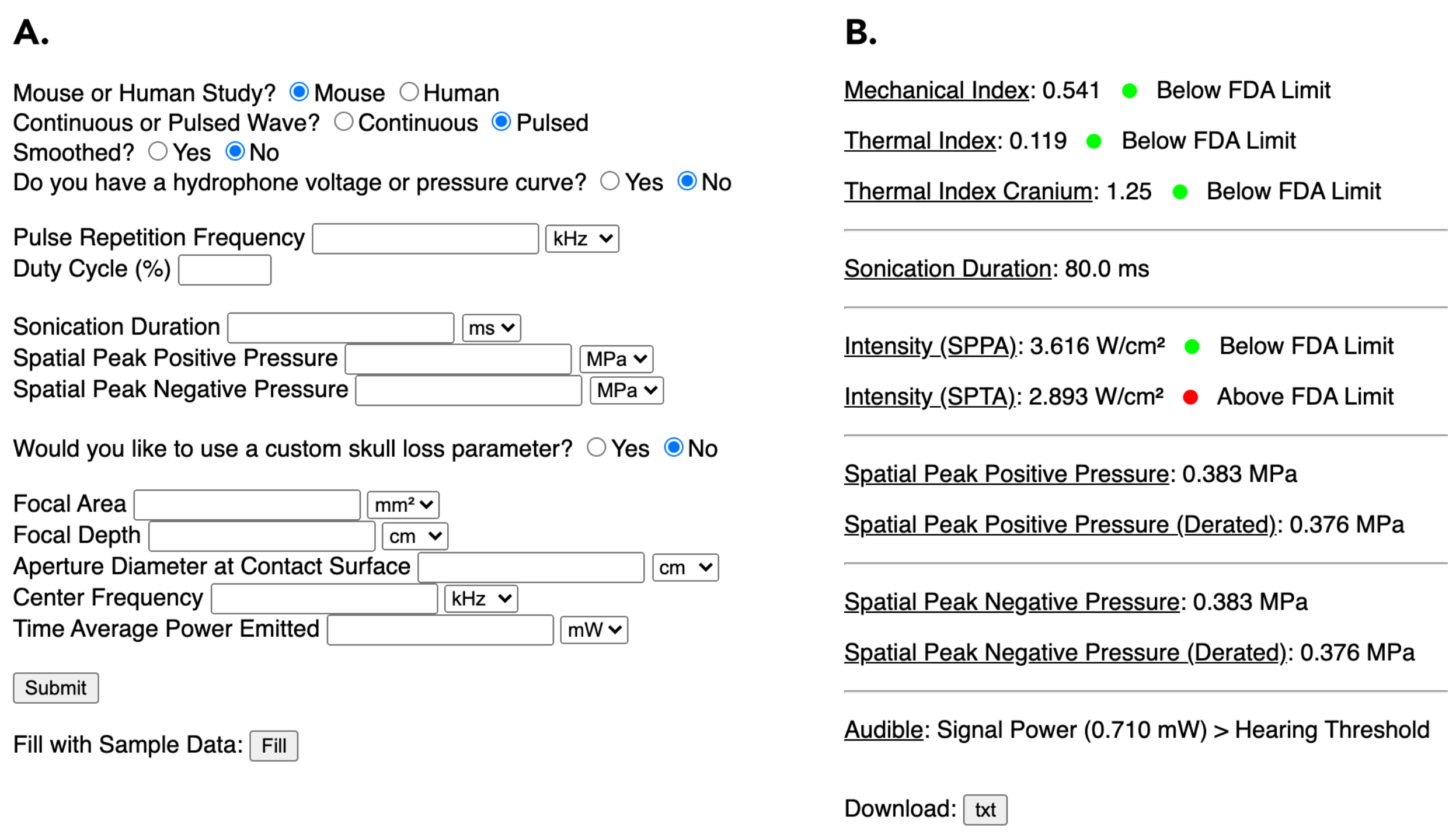

A) Ultrasound parameters inputted by the user. B) Returned safety and wave metrics to be reported by the user in the relevant literature.